Dr. Natacha Gillet
- Postdoc
- room: 819
- phone: +49 721 608 45704
- fax: 45710
- natacha gillet ∂does-not-exist.kit edu
natachagil@orange.fr
Project and CV
Project: electron transfers in Protein and QM/MM simulations
Electrons transfer (ET) processes are very important in biological systems like photosynthesis, cellular respiration, enzymatic process... Understand the role of the protein environment in controlling these efficient mechanisms can lead to new drugs and new materials for energy storage and production. Since decades, computational chemists have developed new approaches to describe charge transfers at the atomic level. Multiscale simulations, in which redox partners are treated at quantum level and environment following classical physical laws, appear as very convenient tools.
Development and comparison between different QM/MM tools
A common issue in computational electron transfer studies consists of the localization of the electron along the pathway and the definition of the different redox states. During my PhD, I used classical MDs coupled to constrained DFT calculations to simulate diabatic states in the framework of Marcus’s theory (http://www.theses.fr/2014PA112159). Another strategy based on the electronic propagation between the different molecules, efficient for fast ETs has been developed and used in Pr. Elstner’s group (http://www.ipc.kit.edu/tcb/104.php). To go farther, I am trying to implement a combination of different approaches, able to describe fast and slow charge transfers, involving tunneling or hopping mechanisms, and even coupled with proton transfers (frequent with organic redox molecules). Furthermore, in collaboration with different other groups, we performed tests and benchmarks of different computational protocols for charge transfers description.
Protein systems
Cryptochrome and Photolyase
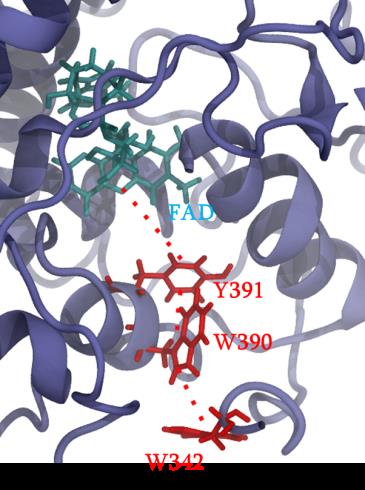
Cryptochrome and Photolyase proteins represent a ubiquitous family of flavoprotein photoreceptors. While the former is involved in plant growth, animal circadian rhythm or perception of the Earth’s magnetic field, the latter participates to DNA repair by oxidizing CPD (cyclobutane pyrimidine dimers) or 6-4 photoproducts. After excitation of a FAD (flavin adenine dinucleotide) cofactor, a fast electron transfer occurs between the protein surface and the flavin through a chain of several aromatic residues. A widely conserved tryptophan triad has been identified but tyrosine or a fourth residue can be sometimes involved in the charge transfer. Some photolyase proteins are currently the object of a strong collaboration with experimentalists. Our study particularly focuses on impact of mutations on the charge transfer kinetics and presence of different charge transfer pathways.
Figure 1 : different partners involves in charge transfer in PhrB protein.
Azurins
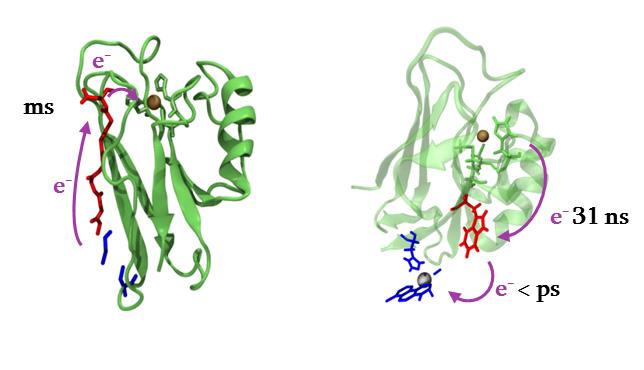
Azurin from Pseudomonas aeruginosa, a type I blue copper protein, has been widely studied as a model for electron tunneling pathway. An extraordinary amount of experiments for biological ETs studies uses this protein, its numerous variants or its dimers. Numerous mutations with natural and unnatural amino-acids have been tested as well as modification of the copper center, or addition of new photoactive metal complex. These data make this protein a good model to test our approaches.
Figure 2 : some electron transfers in (modified) azurins
Ribonucleotide Reductase
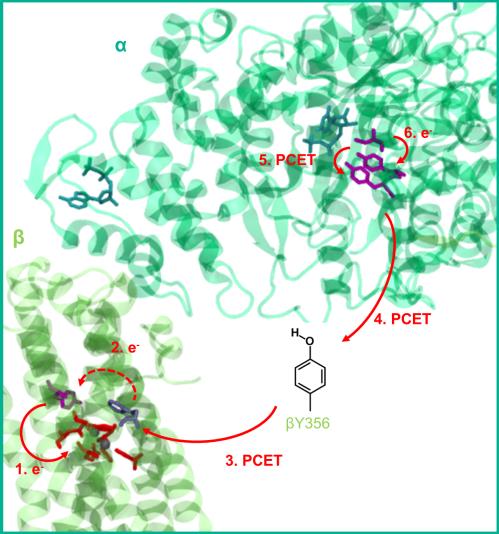
Ribonucleotide reductases (RNR) play a key role in DNA replication and repair by catalyzing the conversion of nucleotides to deoxynucleotides using radical reactions. The class I RNR is form E. coli uses organic radicals generated by a reaction with an iron center. It consists of two homodimeric subunits α2 and β2: α proteins contain the substrate while β subunits contain a diferric tyrosyl radical cofactor. These two centers are separated by about 35 Å. A charge transfer, involving different tyrosine residues takes place when the protein reaches the active conformation. Proton coupled electron transfer simulations are essential to study this systems.
Figure 3 : Proton coupled electron transfer in class I RNR.
Others
I am interested by any collaboration with theoreticians or experimentalists on charge transfer in biological media.
Some science popularization (in French):